|
TRANSFAC - Videos. TRANSFAC (TRANScription FACtor database) is a manually curated database of eukaryotic transcription factors, their genomic binding sites and DNA binding profiles. The contents of the database can be used to predict potential transcription factor binding sites. Since then, the most up- to- date version has to be licensed, whereas older versions are free for non- commercial users. TFs are described with regard to their structural and functional features, extracted from the original scientific literature. They are classified to families, classes and superclasses according to the features of their DNA binding domains. All sites that refer to one TF, or a group of closely related TFs, are aligned and used to construct a position- specific scoring matrix (PSSM), or count matrix. Many matrices of the TRANSFAC matrix library have been constructed by a team of curators, others were taken from scientific publications. Availability. Access to the most up- to- date version requires a license. Applications. The target sequences and the regulated genes can be listed for each TF, which can be used as benchmark for TFBS recognition tools or as training sets for new TFBS recognition algorithms. In the context of systems- biological studies, the TF- target gene relations documented in TRANSFAC were used to construct and analyze transcription regulatory networks. A number of algorithms exist which either use the individual binding sites or the matrix library for this purpose: Patch . Genome Analysis - from Sequence to Function; Bio. Tech. Forum - Advances in Molecular Genetics (J. Database issue): D1. Web Server issue): W2. Web Server issue): W1. Database issue): D1. TRANSFAC's programs, Match and Patch, use the matrices and the site sequences themselves for performing the matrix-or pattern-based search of factor binding sites in regulatory DNA sequences. TRANSFAC and its module TRANSCompel: transcriptional gene regulation in eukaryotes. A new web interface with different search options and integrated versions of Match and Patch provides increased functionality for TRANSFAC. For a better understanding of almost all life processes, a deeper knowledge of gene regulation seems indispensable. The TRANSFAC project as an example of framework technology that supports. Union of Pure and Applied Chemistry (IUPAC) consensus, which can be used for the prediction of TFBSs (PatSearch/Patch. 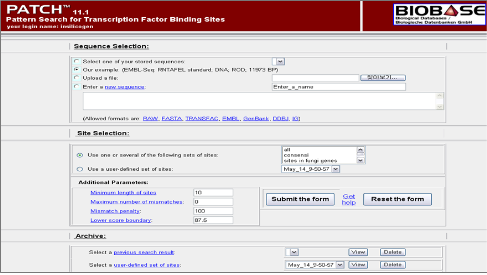
The functionality of Match TM. Match is a weight matrix-based program for predicting transcription factor binding sites (TFBS) in DNA sequences. Learn more about TRANSFAC. Use Patch on this site. P-Match - 1.0 Public. DNA@@@ 5' flanking region. Abalone shriveling syndrome-associated virus. AlloMap molecular expression testing.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. Archives
January 2017
Categories |
 RSS Feed
RSS Feed
